Rare Disease Statistics
Up to 2-6%
of the population worldwide is affected by a rare disease (RD).1-6
Over 7,000
rare diseases have been identified.1,3,6,7
Among rare diseases listed by Orphanet,
~80% are either exclusively genetic or have genetic subtypes.1-3, 8
Half of Rare Disease cases impact children
and 30% of children affected by a RD will not survive beyond the age of 5 years.3,6
The average diagnostic odyssey lasts approximately
5-7 years.
6,9,10
Considering the cost per diagnosis, serial genetic testing in which WGS is used as a last resort test, is
less effective than use as a first-line test.11
For critically ill infants with a Rare Disease, a rapid diagnosis can be critical for timely and appropriate medical intervention.
An early diagnosis can prevent a long, expensive diagnostic journey.12-15
Genetic Testing Approaches for Rare Disease Diagnosis
-
Current standard of care for rare disease may include single gene testing, multi-gene panel testing, chromosomal microarray (CMA) and/or whole-exome sequencing (WES).
-
Whole genome sequencing (WGS) is the only test that can detect nearly all types of genetic variants. (Table 1) 16,17
Table 1
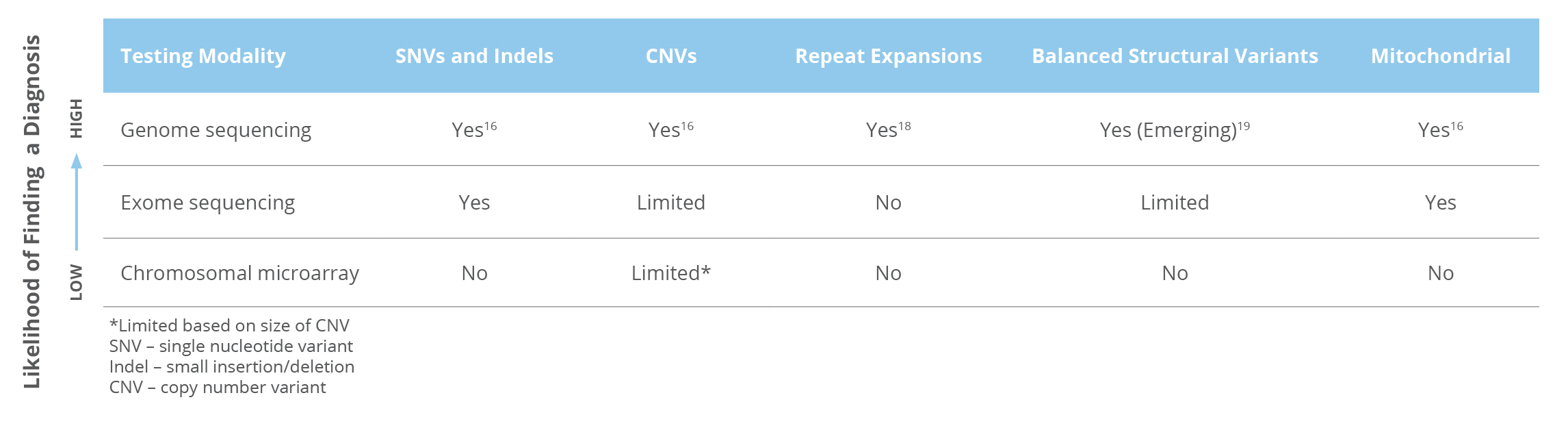
Guideline References
- American College of Medical Genetics and Genomics Board of Directors. Clinical utility of genetic and genomic services: A position statement of the American College of Medical Genetics and Genomics. Genet Med. 2015; 17(6): 505-507.
- Hegde M, Bale S, Bayrak-toydemir P, Gison J, Bone Jeng LJ, Joseph L, Laser J, Lubin IM, Miller CE, Ross LF, Rothberg PG, Tanner AK, VItazka P, Mao R. Reporting Incidental Findings in Genomic Scale Clinical Sequencing d A Clinical Laboratory Perspective A Report of the Association for Molecular Pathology. J Mol Diagnostics. 2015;17(2):107-117. doi:10.1016/j.jmoldx.2014.10.004.
- Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, Grody WW, Hedge M, Lyon E, Spector E, Voelkerding K, Rehm HL, on behalf of the ACMG Laboratory Quality Assurance Committee. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17(5):405-424.
- Matthijs G, Souche E,, Alders M., Corveleyn A, Eck S, Feenstra I, Race V, Sistermans E, Sturm M, Weiss M, Yntema H, Bakker E, Scheffer H, Bauer P. Guidelines for diagnostic next-generation sequencing. Euro J Hum Genet. 2016;24:2–5.
- American College of Medical Genetics and Genomics (ACMG). Policy statement: updated recommendations regarding analysis and reporting of secondary findings in clinical genome-scale sequencing. Genet Med. 2015;17(1):68-69.
- American College of Medical Genetics and Genomics. Points to consider in the clinical application of Genomic Sequencing. a position statement of the American college of medical genetics and genomics. May 2012. epub.
- Boycott K, Hartley T, Adam S, Bernier F, Chong K, Fernandez BA, Friedman JM, Geraghty MT, Hume S, Knoppers BM, Laberge AM, Majewski J, Mendoza-Londono R, Meyn MS, Michaud JL, Nelson TN, Richer J, Sadikovic B, Skidmore DL, Stockley T, Taylor S, van Karnebeek C, Zawati MH, Lauzon J, Armour CM on behalf of the Canadian College of Medical Genetics. The clinical application of genome-wide sequencing for monogenic diseases in Canada: Position Statement of the Canadian College of Medical Geneticists. J Med Genet. 2015;52:431-437.
- Green RC, Berg JS, Grody WW, Kalia SS, Korf BR, Martin CL, McGuire A, Nussbaum RL, O’Daniel JM, Ormond KE, Rehm HL, Watson MS, Williams MS, Biesecker LG. ACMG recommendations for reporting of incidental findings in clinical exome and genome sequencing. Genet Med. 2013;15(7):565-574.
- Kalia SS, Adelman K, Bale SJ, Chung WK, Eng C, Evans JP, Herman GE, Hufnagel SB, Klein TE, Korf BR, McKelvey KD, Ormond KE, Richards CS, Vlangos CN, Watson M, Martin CL, Miller DT on behalf of the ACMG Secondary Findings Maintenance Working Group. Recommendations for reporting of secondary findings in clinical exome and genome sequencing, 2016 update (ACMG SF v2.0): a policy statement of the American College of Medical Genetics and Genomics. Genet Med. 2017;19(2):249-255.
- National Society of Genetic Counselors. Position Statement: Incidental findings in genetic testing. April 2019.
- American College of Medical Genetics and Genomics Board of Directors. Laboratory and clinical genomic data sharing is crucial to improving genetic health care: a position statement of the American College of Medical Genetics and Genomics. Genet Med. 2017;19(7):721-722.
- Position Statement on Licensed Databases and Plans for the Global Sharing of Variant Data. 2015. Accessed February 2019.
- National Society of Genetic Counselors. Clinical data sharing. Position Statement. April 2015.
- Aziz N, Zhao Q, Bry L, Driscoll DK, Funke B, Gibson JS, Grody WW, Hedge MR, Hoeltge GA, Leonard DGB, Merker JD, Nagarajan R, Palicki LA, Robetorye RS, Schrijver I, Weck KE, Voelkerding KV. College of American Pathologists’ laboratory standards for next-generation sequencing clinical tests. Arch Pathol Lab Med. 2014;doi;10.5858/arpa.2014-0250-CP.
- Rehm HL, Bale SJ, Bayrak-Toydemir P, Berg JS, Brown KK, Deignan JL, Friez MJ, Funke BH, Hedge MR. Lyon E, from the Working Group of the American College of Medical Genetics and Genomics Laboratory Quality Assurance Committee. ACMG clinical laboratory standards for next-generation sequencing. Genet Med. 2013;15:733-47.
- Schrijver I, Aziz N, Farkas DH, Furtado M, Ferreira Gonzalez A, Greiner TC, Grody WW, Hambuch T, Kalman L, Kant JA, Klein RD, Leonard DGB, Lubin IM, Mao R, Nagan N, Pratt VM, Sobel ME, Voelkerding KV, Gibson JS. Opportunities and challenges associated with clinical diagnostic genome sequencing: A report of the association for molecular pathology. J Mol Diagn. 2012; 14(6): 525-540.
- Deignan JL, Chung WK, Kearney HM, Monaghan KG, Rehder CW, Chao EC, on behalf of the ACMG Laboratory Quality Assurance Committee. Points to consider in the reevaluation and reanalysis of genomic test results: a statement of the American College of Medical Genetics and Genomics (ACMG). On behalf of the ACMG Laboratory Quality Assurance Committee. Genet Med. 2019;
- American College of Medical Genetics and Genomics Board of Directors. The use of ACMG secondary findings recommendations for general population screening: a policy statement of the American College of Medical Genetics and Genomics (ACMG). Genet Med. 2019; https://doi.org/10.1038/s41436-019-0502-5.
- American College of Medical Genetics and Genomics Board of Directors. Points to consider for informed consent for genome/exome sequencing. Genet Med. 2013; 15(9):748-749.
- Bush, LW, Beck AE, Biesecker LG, Evans JP, Hamosh A, Holm IA, Martin CL, Richards CS, Rehm HL. Professional responsibilities regarding the provision, publication and dissemination of patietn phenotypes in the context of clinical genetic and genomic testing: Points to consider – a statement of the Amercian College of Medical Genetics and Genomics (ACMG). Genet Med. 2018.
- Gibson W, Stavropoulos J, Sinasac D, McCready E, Mahmutoglu S, Nelson TN. Canadian College of Medical Genetics Laboratory Practice Committee. CCMG statement on germline variant classification. 2017.
- Rooney Riggs E, Andersen EF, Cherry AM, Kantarci S, Kearney H, Patel A, Raca G, Ritter DI, South ST, Thorland EC, Pineda-Alvarez D, Aradhya S, Martin CL. Technical standards for the interpretation and reporting of constitutional copy-number variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics (ACMG) and the Clinical Genome Resource (ClinGen).Genet Med. 2019: https://www.nature.com/articles/s41436-019-0686-8
- van El CG, Cornel MC, Borry P, Hastings RJ, Fellmann F, Hodgson SV, Howard HC, Cambon-Thomsen A, Knoppers BM, Meijers-Heijboer H, Scheffer H, Tranebjaerg L, Dondorp W, de Wert GMWR on behalf of the ESHG Public and Professional Policy Committee. Whole genome sequencing in health care. Recommendations of the European Society of Human Genetics. Euro J Hum Genet. 2013;21:580-584.
- Miller DT, Lee K, Gordon AS, Amendola LM, Adelman K, Bale SJ, Chung WK, Gollob MH, Harrison SM, Herman GE, Hershberger RE, Klein TE, McKelvey K, Richards CS, Vlangos CN, Stewart DR, Watson MS, Martin CL & ACMG Secondary Findings Working Group. Recommendations for reporting of secondary findings in clinical exome and genome sequencing, 2021 update: a policy statement of the American College of Medical Genetics and Genomics (ACMG). Genet Med(2021). https://doi.org/10.1038/s41436-021-01171-4
- Miller DT, Lee K, Chung WK, Gordon AS, Herman GE, Klein TE, Stewart DR, Amendola LM, Adelman K, Bale SJ, Gollob MH, Harrison SM, Hershberger RE, McKelvey K, Richards CS, Vlangos CN, Watson MS, Martin CL & ACMG Secondary Findings Working Group. ACMG SF v3.0 list for reporting of secondary findings in clinical exome and genome sequencing: a policy statement of the American College of Medical Genetics and Genomics (ACMG). Genet Med (2021). https://doi.org/10.1038/s41436-021-01172-3
- Rehder C, Bean L.J.H., Bick D,Chao E, Chung W, Das S, O’Daniel J, Rehm H, Shasi V, Vincent LM, ACMG Laboratory Quality Assurance Committee. Next-generation sequencing for constitutional variants in the clinical laboratory, 2021 revision: a technical standard of the American College of Medical Genetics and Genomics (ACMG). Genet Med (2021). https://doi.org/10.1038/s41436-021-01139-4
- Manickam, K., McClain, Demmer LA, et al. Exome and genome sequencing for pediatric patients with congenital anomalies or intellectual disability: an evidence-based clinical guideline of the American College of Medical Genetics and Genomics (ACMG). Genet Med (2021) https://doi.org/10.1038/s41436-021-01242-6
- Souche E, Beltran S, Brosens E, et al. Recommendations for whole-genome sequencing in diagnostics for rare disease. Euro J Hum Genet. 2022: https://doi.org/10.1038/s41431-022-01113-x
Contact Us
Learn more about The Medical Genome Initiative and get answers to your questions.

The Medical Genome Initiative ©2023. All Rights Reserved.
