Overview
Published Manuscript, May 2020
The Medical Genome Initiative: moving whole-genome sequencing for rare disease diagnosis to the clinic 📥
Marshall, C.R., Bick, D., Belmont, J.W. et al. The Medical Genome Initiative: moving whole-genome sequencing for rare disease diagnosis to the clinic. Genome Med. 2020; 12:48. https://doi.org/10.1186/s13073-020-00748-z
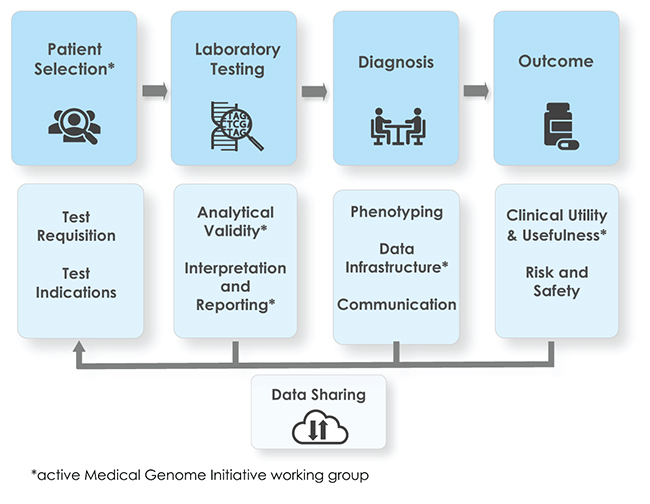
Clinical whole-genome sequencing (WGS) offers clear diagnostic benefits for patients with rare disease. However, there are barriers to its widespread adoption, including a lack of standards for clinical practice. The Medical Genome Initiative consortium was formed to provide practical guidance and support the development of standards for the use of clinical WGS.
Key components of WGS supporting the core steps of genetic disease diagnosis
Analytical Validity
Published Manuscript, October 2020
Analytical validation of clinical whole genome sequencing for germline disease diagnostics: Best practices and performance standards. 📥
Marshall CR, Chowdhury S, Taft RJ, Lebo MS, Buchan JG, Harrison SM, Rowsey R, Klee EW, Liu P, Worthey EA, Jobanputra V, Dimmock D, Kearney HM, Bick D, Kulkarni S, Belmont JW, Stavropoulos DJ, Lennon NJ, on behalf of the Medical Genome Initiative.
Whole-genome sequencing (WGS) has shown promise in becoming a first-tier diagnostic test for patients with rare genetic disorders, however, standards addressing the definition and deployment practice of a best-in-class test are lacking. To address these gaps, the Medical Genome Initiative, a consortium of leading health care and research organizations in the US and Canada, was formed to expand access to high quality clinical WGS by publishing best practices. Here, we present consensus recommendations on clinical WGS analytical validation with a focus on test development, upfront considerations for test design, test validation practices, and metrics to monitor test performance. This work also provides insight into the current state of WGS testing at each member institution, including the utilization of reference and other standards across sites. Importantly, members of this Initiative strongly believe that clinical WGS is an appropriate first-tier test for patients with rare genetic disorders and at minimum is ready to replace chromosomal microarray analysis and whole-exome sequencing. The recommendations presented here should reduce the burden on laboratories introducing WGS into clinical practice and support safe and effective WGS testing for diagnosis of germline disease.
Poster
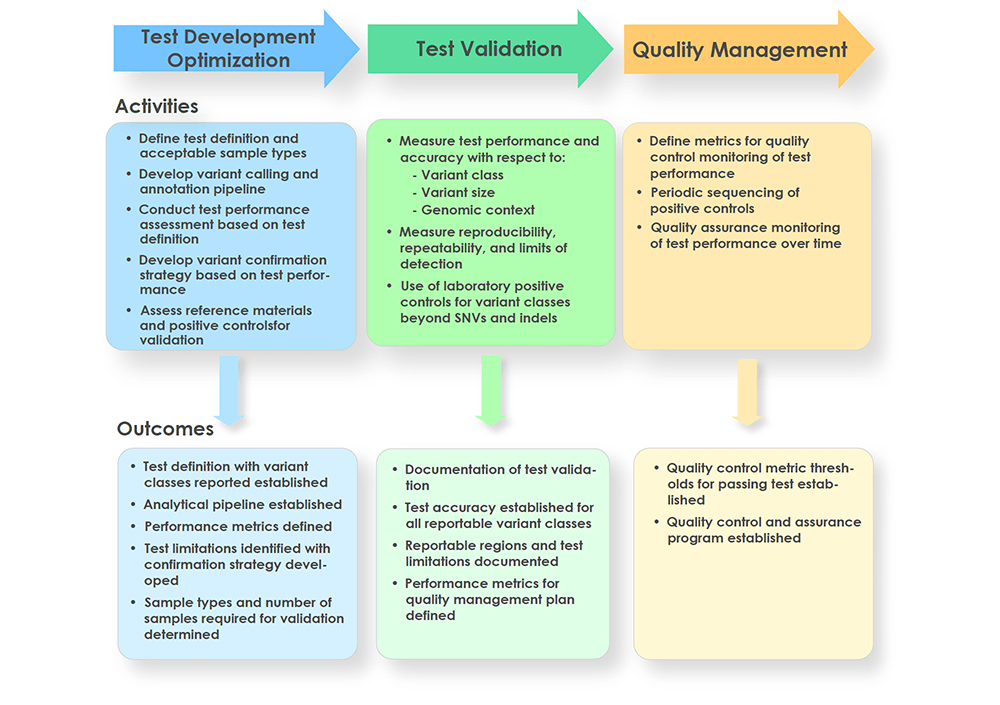
Analytical validation of clinical whole genome sequencing for germline disease diagnostics: Best practices and performance standards. (PDF download) 📥
Marshall CR, Lennon NJ, Chowdhury S, Taft RJ, Stavropoulos DJ, Lebo MS, Harrison SM, Buchan JG, Liu P, Kulkarni S, Dimmock D, Belmont JW, Bick D, Worthey EA, Rowsey R, Klee EW, Kearney HM on behalf of the Medical Genome Initiative. Poster presented at: American Society of Human Genetics Annual Meeting; October, 2019; Houston, TX.
Clinical Utility
Submitted Manuscript
Clinical utility of genomic testing: A measurement toolkit
Hayeems RZ, Dimmock DP, Bick DP, Belmont JW, Green RC, Lanpher B, Jobanputra V, Mendoza R, Kulkarni S, Grove ME, Taylor SL, Ashley E, on behalf of the Medical Genome Initiative.
Whole genome sequencing is emerging as the most robust strategy for achieving timely diagnoses in undiagnosed rare disease populations. Evidence of clinical utility and cost-effectiveness is required for WGS to be accepted into practice, commissioned in a health system, or receive reimbursement. Defining and measuring clinical utility is complex and context specific. The abstract addresses the need to develop a standardized framework and define measurement best practices to optimize the evidence base for decision makers and health care systems invested in providing high quality genome diagnostics.
Manuscript has been submitted to a journal and is under peer review.
Poster
Clinical utility of genomic testing: A measurement toolkit 📥
Hayeems RZ, Dimmock DP, Bick DP, Belmont JW, Green RC, Lanpher B, Jobanputra V, Mendoza R, Kulkarni S, Grove ME, Taylor SL, Ashley E, on behalf of the Medical Genome Initiative. Poster presented at: 2020 American College of Medical Genetics and Genomics Annual Clinical Genetics Meeting; May, 2020; Virtual
*This abstracted was also presented as poster no. P23.08.B at the European Society of Human Genetics meeting held virtually, June 6-9, 2020. The abstract has been accepted as an oral presentation at the European Conference on the Diffusion of Genomic Medicine in spring 2021 (dates to be determined).
Rare Disease Statistics
Up to 2-6%
of the population worldwide is affected by a rare disease (RD).1-6
Over 7,000
rare diseases have been identified.1,3,6,7
Among rare diseases listed by Orphanet,
~80% are either exclusively genetic or have genetic subtypes.1-3, 8
Half of Rare Disease cases impact children
and 30% of children affected by a RD will not survive beyond the age of 5 years.3,6
The average diagnostic odyssey lasts approximately
5-7 years.
6,9,10
Considering the cost per diagnosis, serial genetic testing in which WGS is used as a last resort test, is
less effective than use as a first-line test.11
For critically ill infants with a Rare Disease, a rapid diagnosis can be critical for timely and appropriate medical intervention.
An early diagnosis can prevent a long, expensive diagnostic journey.12-15
Genetic Testing Approaches for Rare Disease Diagnosis
-
Current standard of care for rare disease may include single gene testing, multi-gene panel testing, chromosomal microarray (CMA) and/or whole-exome sequencing (WES).
-
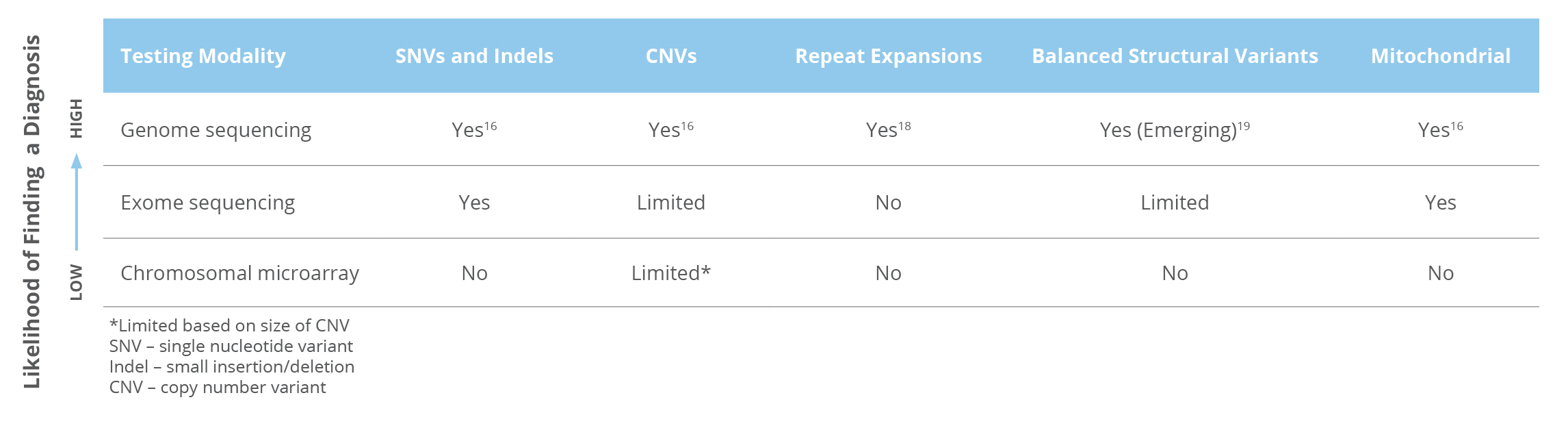
Whole genome sequencing (WGS) is the only test that can detect nearly all types of genetic variants. (Table 1) 16,17
Table 1
References
- Nguengang Wakap, S, Lambert DM, Orly A, et al. Europ J Hum Genet.2019: https://doi.org/10.1038/s41431-019-0508-0.
- Ferreira CR. The Burden of Rare Diseases. American Journal of Medical Genetics. 2019;179(6):885-892.
- Eurodis. About rare diseases. https://www.eurordis.org/content/what-rare-disease. Accessed October 18, 2019.
- Walker et al. The collective impact of rare diseases in Western Australia: an estimate using a population-based cohort. Genet Med. 2017;19(5):546-552.
- Posada De La Paz, M. et al. Rare Diseases Epidemiology: Update and Overview. Advances in Experimental Medicine and Biology. 2017; 1031:589-604.
- Global Genes. Rare disease facts. https://globalgenes.org/rare-diseases-facts-statistics/. Accessed October 18, 2019.
- Online Mendelian Inheritance in Man. https://www.omim.org/statistics/geneMap. Updated October 17, 2019. Accessed October 18, 2019.
- Bick D, Jones M, Taylor SL, et al. Case for genome sequencing in infants and children with rare, undiagnosed or genetic diseases. Journal of Medical Genetics. Published Online First: 25 April 2019. doi: 10.1136/jmedgenet-2019-106111
- Orphanet. About Rare Diseases. https://www.orpha.net/consor/cgi-bin/Education_AboutRareDiseases.php?lng=EN. Updated October 18, 2019. Accessed October 18, 2019.
- Global Commission. Ending the diagnostic odyssey for children with a rare disease. https://www.globalrarediseasecommission.com/Report/assets/static/documents/GlobalCommission-print-021919-a68c8ce2a5.pdf. Published February 19, 2019. Accessed October 18, 2019.
- Gonzaludo N, Belmont JW, Gainullin VG, Taft RJ. Estimating the burden and economic impact of pediatric genetic disease. Gen Med. 2018;
- Soden SE, Saunders CJ, Willig LK, et al. Effectiveness of exome and genome sequencing guided by acuity of illness for diagnosis of neurodevelopmental disorders. Sci Transl Med. 2014;6(265): 265ra168.
- Petrikin JE, Cakici JA, Clark MM, et al. The NSIGHT1-randomized controlled trial: rapid whole-genome sequencing for accelerated etiologic diagnosis in critically ill infants. NPJ Genom Med. 2018 Feb 9;3:6. doi: 10.1038/s41525-018-0045-8.
- Willig LK, Petrikin JE, Smith LD, et al. Whole-genome sequencing for identification of Mendelian disorders in critically ill infants: a retrospective analysis of diagnostic and clinical findings. Lancet Respir Med. 2015;3(5): 377–387. doi:10.1016/S2213-2600(15)00139-3.
- Kingsmore F, Cakici JA, Clark MM et al. A randomized controlled trial of the analytic and diagnostic performance of singleton and trio, rapid genome and exome sequencing in ill infants. A J Hum Gen, 2019;105:1-15.
- Lionel AC, Costain G, Monfared N, et al. Improved diagnostic yield compared with targeted gene sequencing panels suggests a role for whole-genome sequencing as a first-tier genetic test. Genet Med. 2017; Aug 3. doi: 10.1038/gim.2017.119.
- Sanghvi RV,Buhay CJ, Powell, V et al. Characterizing reduced coverage regions through comparison of exome and genome sequencing data across 10 centers. Genet Med. 2017; doi: http://doi.org/10.1038/gim.2017.192.
- Dolzhenko E, van Vugt JJ, Shaw RJ, et al. Detection of long repeat expansions from PCR-free whole-genome sequence data. Genome Res. 2017; 27(11):1895-1903. doi: 10.1101/gr.225672.117
- Chen, X., Schulz-Trieglaff, O., Shaw, R., et al. Manta: Rapid detection of structural variants and indels for germline and cancer sequencing applications. Bioinformatics, 2016;32(8):1220–1222. http://doi.org/10.1093/bioinformatics/btv710
Contact Us
Learn more about The Medical Genome Initiative and get answers to your questions.

The Medical Genome Initiative ©2020. All Rights Reserved.